Like this better. Then one can be explicit about open, close, and the half- variants. Defaults: include_min=include_max=1?
Of course, at least to stay compatible with the previous versions ![]()
+1 that. At least debug(), which is clearly a forensic tool. I’m a bit chary about overly verbose print() output, but this is a matter of taste, perhaps.
While adding this feature to print(), I actually found a bug with references:
$ gmic sp lena eval "(I(256,256) = 1111) = 100;"
does not behave as expected… (only the red component is set to 100).
I’m preparing for another debugging evening! ![]()
Really?
gosgood@bertha ~/git_repositories/gmic-community/tutorial/Beginner_s_Cookbook/Arabesques $ gmic sp lena eval "(I(256,256) = 1111) = 100;"
[gmic]-0./ Start G'MIC interpreter.
[gmic]-1./ Input sample image 'lena' (1 image 512x512x1x3).
[gmic]-1./ Evaluate expression '(I(256,256) = 1111) = 100;' and assign it to status.
[gmic]-1./ Display image [0] = 'lena'.
[0] = 'lena':
size = (512,512,1,3) [3072 Kio of float32].
data = (225,225,223,223,225,225,225,223,225,223,223,223,(...),78,78,78,77,91,80,79,89,77,79,79,82).
min = 8, max = 1111, mean = 128.244, std = 58.972, coords_min = (457,60,0,1), coords_max = (256,256,0,1).
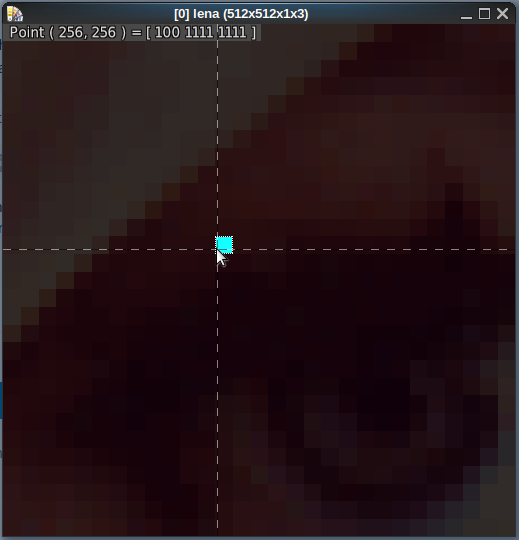
As it should be — no? Or am I entirely misreading this?
gosgood@bertha ~/git_repositories/gmic-community/tutorial/Beginner_s_Cookbook/Arabesques $ gmic -version
gmic: GREYC's Magic for Image Computing: command-line interface
Version 3.2.0 (pre-release #221126)
(https://gmic.eu)
Copyright (c) Since 2008, David Tschumperlé / GREYC / CNRS.
(https://www.greyc.fr)
Built yesterday from the gmic/src develop branch. CImg.h from the current master branch in its repository.
Ah! I am misreading this. The pixel should be [100,100,100].
Exactly. And I found the bug, so I’ll be able to release a bugfix for 3.2.0_pre tomorrow ![]()
does echo_file really appends?
maybe +echo_file could?
Fixed:
New pre-release binaries (3.2.0_pre) now available on the G’MIC website.
I think I found another bug.
Run this to see what I mean:
rep_bug_test:
f "begin(
offset_coords_list=[-1,-1,0,-1,1,-1,1,0,1,1,0,1,-1,1,-1,0];
offset_coords_pos=(offset_coords_list)[1*2,2];
offset_coords_pos*=50;
print(offset_coords_pos);
);
j(offset_coords_pos,0,0,0,2);
"
It’s missing the last two arguments for 0,2.
I found this while solving someone’s problem. This was the solution, but only if vector would be seen as individual arguments.
#@cli rep_local_binary_pattern:
rep_local_binary_pattern:
f "begin(
offset_coords=[-1,-1,0,-1,1,-1,1,0,1,1,0,1,-1,1,-1,0];
);
output_value=0;
repeat(8,pos,
output_value+=(i>j( [(offset_coords)[2*pos,2],0,0,1]))<<pos;
);
output_value;
"
I’m not sure what you mean by this.
j() has two syntaxes:
j(_dx,_dy,_dz,_dc,_interpolation_type,_boundary_conditions)
or
j(P,_interpolation_type,_boundary_conditions)
where P is a vector of dimension 1,2,3 or 4.
And if size(P)<4, the “missing” components are defaulted to 0, so for instance, P = [ -1,-1 ] is equivalent to P = [ -1,-1,0,0 ].
I do not think that was specified in the manual.
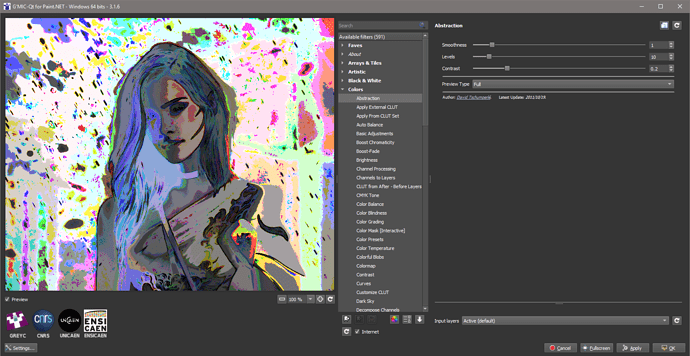
I’m still stuck on 3.16 for GUI filters, but question regardless. Is Abstraction is suppose to look like this?
Also, really cool effect.
I could always read the updatexxx.gmic file in notepad++. Now I try to read the update320.gmic file in Notepad++. Now it shows this:
1 uint8 little_endian
1 5362933 1 1 #1481633
xœì½i»ÛDÖ(úýü
߼Ͻ7y½M¶eseÎñ<HžaYî·oY’-Ù²$kðÔÏyžì„! „ ÍІI„С› >ô?¹;Þ{ç_Ü’d{{Ü{aú=çM Ž]µªT¥Zµ¦ZkÕ¿ýfGdÿÛ¿ý·s¹b¢Ä»¦žr™*Ç<†Âg×òŸ“®øÿN†]¬Òé02çjXíO9}ExÕDÕÙê«l÷å@4m±¡îš´6xMß²ë{à›Õ;ƒž]'U?ñÏè¼ëßPAÐÀ©SgÖI0U
What am I doing wrong? How can I convert this to something readable? Or where can I find the readable version?
How did you get this update320.gmic file?
If it’s from https://gmic.eu/update320.gmic, then yes, it is stored in a compressed binary form.
The non-compressed version is here: https://gmic.eu/plain_update320.gmic
When you push the ‘Update’ button from the G’MIC-Qt plug-in, it should store this file as an uncompressed ASCII file in your %APPDATA%\gmic folder (named update320.gmic).
I did a new instal of gimp and gmic. The update button first didn’t work. I had to update via the properties menu. After that, the “usual” update button works also. And yes now I can read the file, thank you David.
by drawing perpendicular hairs to the edge
i was surprised by the swap of vectors between structure and diffusion tensors. i don’t think it’s a bug, and i can easy add line borrowed from the original.
(structureT is alike and quicker then diffT)
ee:
eigen s. c
v 1 e =$1=cos={i(#-2)}===sin={i(#-1)}==={acos(i(#-2))/pi*180}°== v -
foo:
10,10 f. ‘y’
+structuretensors[0] , ee. ST
+diffusiontensors[0] 0,1,0,0 ee. DT
+l[0] { g xy,0 +* sqr[0,1] compose_channels + rv[1,2] a c } ee. S0
+l[0] { g yx,0 +* sqr[0,1] compose_channels + rv[1,2] a c } ee. S1
=ST=cos=0===sin=1===90°==
=DT=cos=1===sin=0===0°==
=S0=cos=0===sin=1===90°==
=S1=cos=1===sin=0===0°==
Yes, basically, eigenvectors of structure tensors and diffusion tensors are the same, but the order of the associated eigenvalues are reverted.
Reporting a possible bug now because I’ll forget otherwise… command orthogonalize appears to do nothing. I think due to the arguments list provided to draw().